Comments (27)
MPIBackend and JoblibBackend now provide unique seed states for each trial in a simulation that is consistent across backends with #171. What is left to do here is
- Validate results with HNN
- Refactor the process (including updating the API) for how seed states are assigned
from hnn-core.
Right now, MPIBackend doesn't set unique seeds for each trial. That should be fixed in a PR that addresses this issue. Also JoblibBackend has a scheme that could be improved.
https://github.com/jonescompneurolab/hnn-core/blob/master/hnn_core/parallel_backends.py#L88-L90
This doesn't guarantee unique seeds for each trial. For example, if a prng_ seed is set to 1, then the first two trials will have the same seed.
HNN's scheme of setting the seed is perhaps unique, but very convoluted. The function reset_src_event_times was adapted to hnn-core in #55:
https://github.com/jonescompneurolab/hnn-core/pull/55/files/39c6f8800fa5e28ecf785cda1ef5440626833c43#diff-535c9c5d7e5fe03d2cd31f20bd77fc25R263
It had different behavior for incrementing the seed depending on if an input is part of Network.extinput_list (increment by 1000)
https://github.com/jonescompneurolab/hnn-core/pull/55/files/39c6f8800fa5e28ecf785cda1ef5440626833c43#diff-535c9c5d7e5fe03d2cd31f20bd77fc25R270
or part of Network.ext_list (increment by 1):
https://github.com/jonescompneurolab/hnn-core/pull/55/files/39c6f8800fa5e28ecf785cda1ef5440626833c43#diff-535c9c5d7e5fe03d2cd31f20bd77fc25R277
@jasmainak what about changing
if trial_idx != 0:
net.params['prng_*'] = trial_idx
to
net.params['prng_*'] += trial_idx
Can we implement an HNN-backward compatibility inc_prng function that will be deprecated once tests are done comparing to HNN and #136 is closed? See
https://github.com/jonescompneurolab/hnn-core/pull/55/files/39c6f8800fa5e28ecf785cda1ef5440626833c43#diff-91f1465772612a9e40c10b7dfab6c5f6R41
from hnn-core.
That's fine changing to:
net.params['prng_*'] += trial_idxBy the way, I think I finally understood why this increment of 1000 is done. It's because for the unique input feeds, you have a separate random number generator for each cell, so you add 1000 to avoid the possibility of overlap?
In my opinion, we should try to use the numpy interface for random number generation as much as possible rather than developing an ad hoc procedure which is going to be prone to bugs. For e.g., you can generate a series of 2D floats and then send them to appropriate processes using either backend. This way we keep the randomness in the codepath outside of parallel.
from hnn-core.
In my opinion, we should try to use the numpy interface for random number generation as much as possible rather than developing an ad hoc procedure which is going to be prone to bugs. For e.g., you can generate a series of 2D floats and then send them to appropriate processes using either backend. This way we keep the randomness in the codepath outside of parallel.
Sounds like a good plan. As long as there is a HNN-compatible method too that will be deprecated.
from hnn-core.
That's fine changing to:
net.params['prng_*'] += trial_idxBy the way, I think I finally understood why this increment of 1000 is done. It's because for the unique input feeds, you have a separate random number generator for each cell, so you add 1000 to avoid the possibility of overlap?
In my opinion, we should try to use the numpy interface for random number generation as much as possible rather than developing an ad hoc procedure which is going to be prone to bugs. For e.g., you can generate a series of 2D floats and then send them to appropriate processes using either backend. This way we keep the randomness in the codepath outside of parallel.
So the plan down the road (i.e., after we've created a HNN-compatible method) is to have a system where we are no longer incrementally change net.params['prng_*'] right? For clarity's sake, I'm imagining that hnn-core will contain a one-time assignment that 1) instantiates the np.random.RandomState container and then 2) generates all jittered feed start times across trials which can then be passed to parallelized trials/processes. Then, we can add an argument to the ExtFeed constructor that allows the user to supply a single seed if they wish, otherwise, np.random.RandomState() will use a default seed. This will also eliminate the need to specify any 'prng_*' parameters in the param files. @blakecaldwell @jasmainak is this what you both had in mind?
from hnn-core.
yes, exactly that is the idea.
from hnn-core.
@cjayb you might want to look at this issue and the discussion. I think it's related to the feeds PR. Perhaps with @rythorpe you guys could come up with a plan how to move forward on this? Please take a look at this to understand what the GUI currently does.
from hnn-core.
I wanted to check on the status of this. Is anything else needed @cjayb @rythorpe?
from hnn-core.
We added this property
https://github.com/jonescompneurolab/hnn-core/blob/master/hnn_core/network_builder.py#L268
in order to enable this indexing
https://github.com/jonescompneurolab/hnn-core/blob/master/hnn_core/network_builder.py#L402
that stems from the new 'feed instantiation' moved to Network:
https://github.com/jonescompneurolab/hnn-core/blob/master/hnn_core/network.py#L315
I had a brief struggle with one of the examples failing to produce different dynamics for consecutive trials here
https://github.com/jonescompneurolab/hnn-core/blob/master/examples/plot_simulate_evoked.py#L49
but fixed that in dipole.py.
That's a long way of saying: I'm pretty sure the trial seeding is reproducible now.
from hnn-core.
That said, I totally second @rythorpe and @jasmainak above regarding not repeatedly re-seeding the prng! It's a relic from pre- #191 times, where event times were generated in NetworkBuilder, i.e., potentially on different hosts/ranks. Now everything is done in Network, so there's no reason to do that, other than backward-compatibility with the existing examples...
I'm sketching out a new example to demonstrate a new API for creating the feeds/drives we've been discussing for a while now. Being able to re-create the funky HNN-Classic seeding is causing more work than the API itself :)
from hnn-core.
@blakecaldwell I'm pretty sure, as @cjayb said, that hnn-core now has a fairly stable and consistent method for providing unique seeding across multiple trials. I've been running a few simulations to test the consistent between hnn-gui and hnn-core but something is off. I have yet to determine if the issue is caused by a seeding discrepancy or differences in some other feature.
from hnn-core.
was there an old commit where there was no discrepancy? If so, you can try to run "git bisect" ... I've found it very useful in narrowing things down in the past
from hnn-core.
was there an old commit where there was no discrepancy? If so, you can try to run "git bisect" ... I've found it very useful in narrowing things down in the past
I know we can simulate evoked inputs with no discrepancy, but the problems I'm encountering are when rhythmic inputs are added. I'm not aware of a time when we tested such a case and multiple trials were added.
from hnn-core.
Wild guess: could the cellular targets of the 'common' rhythmic feeds differ? Not just cell type, but location (soma vs dendrites)?
from hnn-core.
single trial works I believe (I checked this at least visually at some point). So it's got to have something to do with seeds. There is an increment of 1000 to look for somewhere.
from hnn-core.
There is an increment of 1000 to look for somewhere.
@jasmainak correct me if I'm wrong, but I think this only applies to incremented evoked inputs - a rarely used feature in hnn that we don't plan on implementing in hnn-core.
from hnn-core.
This issue is resolved upon validating that the hnn-core output is consistent (explainable difference permissible) with that of HNN for all the following drives. (Please specify whether a single or multiple trials were validated and link to commits to hnn-core, and hnn or a tutorial image in hnn-tutorials.)
- evoked drives as defined in
default.jsonproduce multiple identical trials in ea2f3a7 and jonescompneurolab/hnn@a09bb08) - bursty (rhythmic) drives as defined in the alpha example at 58efb61 match the results of the hnn tutorial at jonescompneurolab/hnn-tutorials@12fcea9 (@ntolley)
- poisson drives (@cjayb @jasmainak ) any commit before jonescompneurolab/hnn@90d992a and any commit before beb590f should match
-
tonic baseline(tonic baselines are implemented as constant cellular mechanisms and are independent of rng seeding)
from hnn-core.
I can look into this after Feb 12 :)
from hnn-core.
I can look into this after Feb 12 :)
Totally fine @jasmainak. I'll probably be able to resolve some of these on my own this week - just wanted to keep y'all in the loop and request help if any of the items are quick fixes.
from hnn-core.
Running make doc on 58efb61 produces the following results with reference to HNN GUI documentation
Alpha example (bursty/rhythmic drives)
Compare example output to relevant portion of Figure 7.
- I'd say this is a "check'
Gamma example (Poisson drive and tonic current clamp)
There are inconsistencies between HNN GUI output (current master) and the documentation. These need to be resolved before hnn-core output can checked against a standard.
from hnn-core.
Gamma example (Poisson drive and tonic current clamp)
There are inconsistencies between HNN GUI output (current master) and the documentation. These need to be resolved before
hnn-coreoutput can checked against a standard.
Are you referring to jonescompneurolab/hnn#264 or are there other discrepancies as well?
from hnn-core.
Using the L5/L2 weak param file doesn't reproduce the Tutorial figure 4, I think, in current GUI master. Can you (de)confirm?
from hnn-core.
In my tests, HNN GUI (master) does not reproduce the website tutorial. I'm guessing it has to do with L89 in feed.py:
t_gen = t0 - lamtha * 2 # start before t0 to remove artifact of lower event rate at start of periodIf I change that to
t_gen = t0and run HNN GUI master with gamma_L5weak_L2weak.param, I get
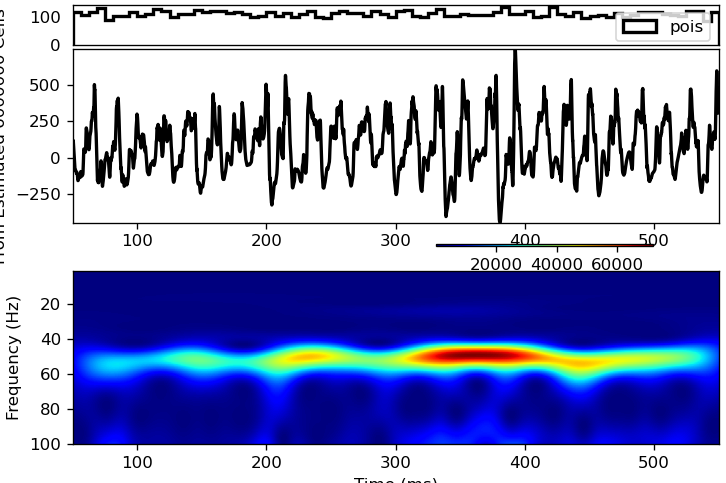
I can reproduce this with hnn-core master, using the new API and setting seedcore to 1079 (which, confusingly, logically ought to correspond to prng_seedcore_extpois=-3...)
Note that there's a difference between gamma_L5weak_L2weak.param and gamma_L5weak_L2weak.json (hnn-core) in the seedcore for the Poisson process.
Take home message(?)
hnn-core can reproduce GUI output for Poisson drives, but there's something wonky going on wrt. seed assignments. I would say, though, that this is neat yet another
- check
from hnn-core.
hnn-corecan reproduce GUI output for Poisson drives, but there's something wonky going on wrt. seed assignments. I would say, though, that this isneatyet another
- check
Thanks for looking into this @cjayb. I had this on my to-do list but you beat me to it. I'm assuming that the difference in how hnn-core and HNN treat the seedcore param has to do with how we set gids?
from hnn-core.
Sure.
difference in how hnn-core and HNN treat the
seedcoreparam has to do with how we set gids?
Well, it's a combination of params-files sometimes using negative seedcore-values, and the fact that HNN GUI always adds certain drive (feed) gids, regardless of whether they are used. In the end, it turns out that we need to set the Poisson seed to 270 + 2 + 3 x 320 - 3 = 1079... Long story...
from hnn-core.
I edited the description above for the Poisson. Didn't check it again now but I think it's correct. Please feel free to cross-verify @rythorpe. Note that prior to these two commits, there was a bug in both HNN and HNN-core because np.append return value was not used. While in HNN-core, this was fixed in beb590f, in HNN an additional change was made to account for the burn-in.
from hnn-core.
Discrepancies in seeding between HNN and hnn-core have been removed or accounted for in each drive/bias type as outlined in #113 (comment).
from hnn-core.
Related Issues (20)
- BUG: MPI simulations break for `dt` of certain size HOT 1
- Upload data error - HNN gui HOT 2
- Clean up optimization API
- Improve error messages for adding drives
- rename simulate_dipole -> simulate HOT 3
- Proposed Enhancements for the Current GUI: A Refined Feature List HOT 1
- change name from calcium model to Kohl_2022 HOT 4
- [JOSS Review] HNN-core: A Python software for cellular and circuit-level interpretation of human MEG/EEG HOT 2
- [JOSS] Documentation HOT 1
- Optimization example error HOT 4
- Problems using non-'soma' values for record_isec argument in simulate_dipole() HOT 12
- [JOSS] Software Paper HOT 2
- pre-allocate arrays for storing continuous simulation data in network_builder.py
- issue with GUI install
- GUI does not show dipole plots HOT 2
- installation on m2 mac HOT 1
- [BUG] `plot_dipole` not showing in GUI with `matplotlib>=3.8.0` HOT 9
- tests: add axis data checks for all plots available in GUI HOT 2
- BUG: Deleting drives in GUI after file upload prevents loading of the same file HOT 1
- GUI callbacks error messages are not logged
Recommend Projects
-
 React
React
A declarative, efficient, and flexible JavaScript library for building user interfaces.
-
Vue.js
🖖 Vue.js is a progressive, incrementally-adoptable JavaScript framework for building UI on the web.
-
 Typescript
Typescript
TypeScript is a superset of JavaScript that compiles to clean JavaScript output.
-
TensorFlow
An Open Source Machine Learning Framework for Everyone
-
Django
The Web framework for perfectionists with deadlines.
-
Laravel
A PHP framework for web artisans
-
D3
Bring data to life with SVG, Canvas and HTML. 📊📈🎉
-
Recommend Topics
-
javascript
JavaScript (JS) is a lightweight interpreted programming language with first-class functions.
-
web
Some thing interesting about web. New door for the world.
-
server
A server is a program made to process requests and deliver data to clients.
-
Machine learning
Machine learning is a way of modeling and interpreting data that allows a piece of software to respond intelligently.
-
Visualization
Some thing interesting about visualization, use data art
-
Game
Some thing interesting about game, make everyone happy.
Recommend Org
-
Facebook
We are working to build community through open source technology. NB: members must have two-factor auth.
-
Microsoft
Open source projects and samples from Microsoft.
-
Google
Google ❤️ Open Source for everyone.
-
Alibaba
Alibaba Open Source for everyone
-
D3
Data-Driven Documents codes.
-
Tencent
China tencent open source team.




from hnn-core.